Monitoring Bioreactors With In-Situ NIR
In the past decade, near-infrared (NIR) spectroscopic analysis has become an important analytical tool for real-time, in-situ monitoring of bioprocesses—in the spirit of the FDA’s process analytical technology (PAT) initiative. In order to use NIR successfully for bioprocess applications, robust prediction models must be developed, which requires an understanding of key features of the technology and how it is best employed. This article discusses those features and describes a unified approach to robust model development and the successful implementation of NIR analysis of bioprocesses.
Slow to Take
A fundamental element of PAT is the use of in-line analysis to increase process understanding and control and to verify product quality prior to release. For bioprocesses, in-line monitoring using NIR has been applied for years [1], yet its rate of implementation has been slower than expected. This is due in part to the relative lack of professional NIR practitioners compared with those for HPLC, mid IR and other technologies. Implementation has also been hindered by the biopharmaceutical industry’s proprietary environment, which often forces NIR practitioners to work in isolation. Few biopharma companies have published anything but general discussions of NIR applications. An exception has been the Strathclyde Fermentation Center at the University of Strathclyde in Glasgow, U.K. [2].
Analyzing bioprocesses in-line is always a challenge. The complexity of the bioreactor media creates a matrix of absorbance bands. Often, commercial media may derive from a proprietary recipe, and a media matrix may contain literally hundreds of nutrients and buffers. Nutrients are depleted throughout the run, but some are fed back into the bioreactor when nutrient levels become critically low. In addition, metabolites like lactate and ammonia increase with cell growth throughout the run and must be neutralized to reduce their concentration below levels that would be toxic to the cells. The cells do increase through the run, but not geometrically like microbial cells and therefore transmission spectroscopy can be used. The titer (hopefully) increases throughout the run.
With NIR in-situ monitoring, these process attributes can be monitored and controlled with feedback (or feed forward) through digital or analog I/O connectivity to the bioreactor PLC.
Some background as to why: the fundamental absorption bands of chemical functional groups occur in the mid-infrared region of the electromagnetic spectrum. These absorptions are very strong and dilutions, very short pathlengths or ATR (attenuated total reflectance) methods are required to bring the absorbances within the linear range of the detector. The overtone absorptions of these fundamental bands occur in the NIR spectral region, which is between the mid-IR and the visible. This region allows direct measurement without sample preparation due to the relative weakness of absorption.
Since NIR sources are very powerful and detectors for this region of the spectrum
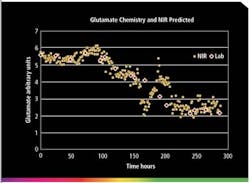
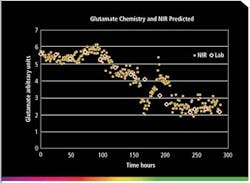
Figure 1. Trend plot of glutamine level in a 100-liter bioreactor over 12 days. A feed occurred after 150 hours.
are very sensitive, measurements with high signal-to-noise are possible. Economic fiber-optic cables can be used to measure processes at locations remote from the analyzer through the use of probes inserted into bioprocessing equipment. With fused silica fibers that do not absorb strongly in the NIR region, one instrument can be multiplexed to monitor up to nine bioreactors.
Challenges to Model Development
Common analytes that have been measured with NIR in situ in bioreactors include: glucose, lactate, glutamine, glutamate and ammonia. Amino acid levels and product titer have also been measured, as have bioprocess parameters including pH and cell density [3].
However, NIR is very sensitive to analyte levels in the complex matrix of the growth media. Thus, there can be problems with calibration model development:
- The absorption bands overlap in NIR, so variance of one analyte can change the prediction of others. Therefore, all variance from expected analytes (within range of real operation) must be included for PLS modeling.
- Also, “non-analyte” variance (that is, variance from sources not commonly thought of as chemical species) must be considered in order to develop a robust model that will predict well on all subsequent runs. These non-analyte variables may include: temperature; pH; probe-to-probe variance; channel-to-channel variance; instrument-to-instrument optical variance; type and manufacture of stock growth media; cell type or line; and titer molecule produced.
This variance is not completely contained in a single two-week bioreactor run, and thus multiple runs are necessary. The initial goal is to make a simple model for a single media type with a single cell type and a single titer. The models should later be developed for multiple probes that may initially have bias and slope changes. The next goal would be to develop more global models that can predict on various media types and cell types and titer.
Reference data for a given bioreactor are usually collected only once per day. Some analyte values are available rapidly, such as those for glucose, lactate, ammonia, glutamine and glutamate from at-line blood analyzers (such as the NOVA BioProfile Analyzer). Data for the other amino acids (and more accurate glutamine and glutamate values) are sought via HPLC and are typically not available for many days or weeks.
The environment in the reactor due to pH and temperature is kept very stable, but there may be a step change at initial inoculation or infection. The pre-inoculation period usually has very little reference data for modeling, so in order to model it well, data from this period, accounting for several runs, must be used.
NIR spectra can be collected every minute or less frequently, if desired. This leads to the accumulation of a tremendous amount of data that should be archived daily. Often half-hour or hourly data is sufficient to understand trends over a two-week period. Figure 1 shows a trend plot of glutamine level taken hourly in a 100-liter bioreactor over twelve days. A feed occurred after 150 hours.
The Case for Chemometrics and NIR Modeling
Chemometrics for NIR analysis uses mathematical and statistical methods and algorithms to solve spectrochemical problems. This includes mathematical pretreatments of the spectra to enhance linear chemical variance, regression analysis methods and modeling diagnostics.
The first thing to do in modeling bioreactor spectra is to normalize the baseline with a math pretreatment like second derivative and standard normal variate (SNV) or multiplicative scatter correction (MSC). This removes the scattering effects of the cells and allows chemical absorbances to stand out.
Regions resulting from water must be removed from the analysis, for example, areas where absorbance exceeds the linear range of the detector/probe combination at the pathlength used. If the spectra are well referenced and contain accurate data at the correlated time, a partial least squares (PLS) result with several factors can be obtained. The predicted residual error sum of squares (PRESS) may reach a minimum after several factors or so, and that is a limit for that data set.
With low noise spectra and enough samples and degrees of freedom, it is possible that many factors are justified. This also depends on the concentration of the analyte and the accuracy of the reference data and the exclusion of any “bad” spectra. Bad spectra may be due to bubbles in the gap at the time the spectrum was taken. To optimize the results for that specific analyte, it is best to repeat the PLS modeling for each analyte of interest with slight modifications.
Developing Robust Models
A common approach in bioreactor modeling is to develop an off-line cell-free set of experiments to understand where the analytes will absorb in the media. Some degree of understanding can be gleaned from this method, but the results will not be closely related to the actual media with cells. This is because of the broad band overlapping absorbances, the scattering effects and added absorbance of the cells. The regions that showed high correlated variance in the loading plots in the cell-free media will not necessarily be the same regions in the media with cells.
The factor loading plots in PLS modeling are an important chemometric diagnostic tool. They show the regions of highly correlated spectral absorbance to analyte value and should be checked for uniqueness for each analyte. They can also be used to check that the model is not tracking the inverse of the analyte concentration (as in water reduction due to analyte increase).
This is a starting point for developing a robust model. A model can be built on one cell culture run and used to predict the next. Then, cell culture runs can be combined to build a more robust model and predict real time on successive runs. Several models can be evaluated at once with slightly different modeling and different numbers of factors to see which model works best on the next run.
Detrend polynomial fit can be applied first before the second derivative and SNV. This math treatment may help predict on subsequent runs as it further reduces baseline offset from run to run and probe to probe. These are typical problems experienced with robustness in multi-run/multi-probe applications. It is relatively easy to develop a model for a single run with well correlated and accurate reference data. Developing models that are robust on subsequent runs and multiple probes is more difficult.
The best approach is to start with reasonable variance on one system of media, cell type and titer. Be sure to include all acceptable variance of pH, temperature, cell density and levels of all analytes. An analyte that has little or no change throughout a run will offer no variance within the run, although it may offer useful information when combined with other runs.
There is no sense in including levels of analytes at which the cells would be dead. If the cells are dead, other means of analysis are available! The utility of in-situ NIR is the real time trending of growth limiting nutrients, metabolites of concern (product and waste) and operational parameters such as pH.
Conclusion
NIR in-situ monitoring of bioreactors is a good application of one of the principal tools in the PAT toolbox and can help increase bioprocess understanding and control. The complexity of the bioreactor media creates a matrix of absorbance bands and a systematic approach to model development is required. It is best to start initially with a simple model for a single media type with a single cell type and a single titer. The necessary variance is not completely contained in one two-week bioreactor run and multiple runs are necessary. The spectra with reference data can be added to the model from the previous run after each new run until all necessary and required variance has been covered in the model. Probe-to-probe variation should be modeled in after the basic chemical and operational parameters have been sufficiently modeled.
References
1. Arnold, S.A., Crowley, J., Woods, N., Harvey, L.M., McNeil, B. In-Situ Near Infrared Spectroscopy to Monitor Key Analytes in Mammalian Cell Cultivation. Biotechnology and Bioengineering. 84(1): 13-19 (2003).
2. Roychoudhury, P., O'Kennedy, R. McNeil, B., Harvey, L.M. Multiplexing fibre optic near infrared (NIR) spectroscopy as an emerging technology to monitor industrial bioprocesses. Anal Chim Acta. 590(1): 110-117 (2007).
3. Mattes, R., Root, D., Chang, D., Ong, M., Molony, M. In Situ Monitoring of CHO Cell Culture Medium Using Near-Infrared Spectroscopy, BioProcess International. 5(01): S46-S51 (2007).